7th MACSMIN 2026: Mathematics and Computer Science for Materials Innovation
 |
The MACSMIN logo includes the basic examples of the rock-salt cubic crystal, the benzene ring, and a blue wave containing a local maximum and a local minimum. |
 |
The CCP-QC (UK Collaborative Computational Project in Quantum Computing) has generously funded travel and accommodation for participants of the 7th MACSMIN. |
- Dates: 26-29 May 2026 in a hybrid form in the Materials Innovation Factory, Liverpool, UK.
- Organisers: Data Science group including Vitaliy Kurlin, Olga Anosova, and Dan Widdowson.
- The registration is free: e-mail Vitaliy Kurlin, see the the travel information for Liverpool (UK).
- Talks on advances in Geometric Data Science will be given by Daniel Widdowson, Olga Anosova, Yury Elkin, Ziqiu Jiang, Surya Majumder, Jack Gallimore, Gabriel Newton, and Vitaliy Kurlin.
- All talks will be aimed for a broad audience of scientists including PhD students and postdocs.
- Related meetings: mini-symposium Computational Data Science of Nanostructures (6 hours) at the SIAM annual meeting on July 28 - August 1, 2025 in Montreal: part 1, part 2, part 3.
- The AMS session on Open Problems in Geometric Data Science at JMM 2026, Washington DC.
- The ICERM workshop Rigidity Theory meets Geometric Data Science for applications in chemistry and biology on 6-10 July 2026, Brown University, Providence (US).
- The regular MIF++ seminar is a continuous version of the annual MACSMIN.
- Past meetings of the MACSMIN series: 2025, 2024, 2023, 2022, 2021, 2020.
Back to Top of this page | Back to MACSMIN | Back to Home page
 From left to right:
Saulius Gražulis,
Alexandre de Brevern,
Thérèse Malliavin,
Richard Catlow FRS,
Wolfgang Hornfeck,
Alex Wlodawer,
Olga Anosova,
Paweł Rubach,
Dan Widdowson,
Vitaliy Kurlin,
Aaron Finney, and online participants above.
From left to right:
Saulius Gražulis,
Alexandre de Brevern,
Thérèse Malliavin,
Richard Catlow FRS,
Wolfgang Hornfeck,
Alex Wlodawer,
Olga Anosova,
Paweł Rubach,
Dan Widdowson,
Vitaliy Kurlin,
Aaron Finney, and online participants above.
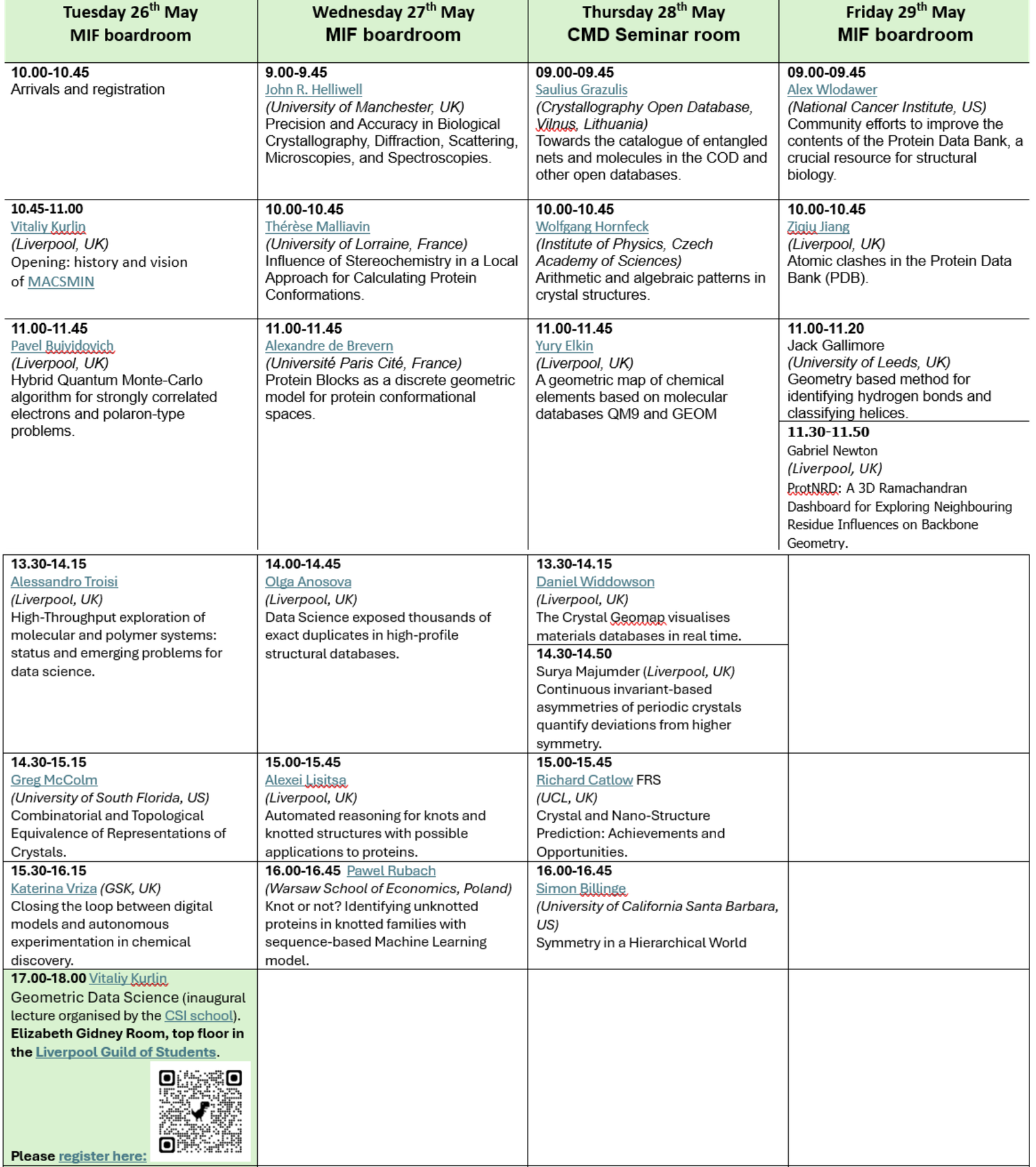
Timetable (all UK times): pdf Tuesday Wednesday Thursday Friday
Tuesday 26 May (the MIF boardroom)
- 10.45-11.00 (in person and on zoom)
- Opening: history and vision of MACSMIN by Vitaliy Kurlin (Liverpool MIF, UK)
- 11.00-11.45 (in person and on zoom)
- Pavel Buividovich (Mathematical Sciences, University of Liverpool, UK)
- Title. Hybrid Quantum Monte-Carlo algorithm for strongly correlated electrons and polaron-type problems.
- Abstract. We discuss quantum Monte-Carlo simulations, in particular, the Hybrid Monte-Carlo (HMC) algorithm as a tool to study charge transport in quantum materials. Traditional applications of QMC/HMC include studies of spontaneously ordered phases and quantum phase transitions at low temperatures. We discuss a less traditional application of HMC to study charge transport in molecular organic semiconductors at high temperatures. We outline some conceptual challenges of studying real-time charge transport using QMC/HMC, including the ill-defined numerical analytic continuation from imaginary to real time. We then discuss several recent improvements of the HMC algorithm that allow to significantly enhance its precision and hence its predictive power for real-time responses such as electric conductivity. We demonstrate that, in a less traditional formulation, HMC is also well suited to study polaron-type problems where a single charge carrier interacts with a high-temperature phonon bath.
- 12.00-13.30 lunch (covered for speakers by the organisers)
- 13.30-14.15 (in person and on zoom)
- Alessandro Troisi (Chemistry, University of Liverpool, UK)
- Title. High-Throughput exploration of molecular and polymer systems: status and emerging problems for data science.
- Abstract. This lecture will review two materials areas explored in our group using computational chemistry methods highlighting the success but also the requirement of novel methodology to fully exploit and develop such datasets. A first area is the exploration of optical and electronic properties of single molecules. There is a natural clustering suggested by chemical intuition that allows one to explore virtually the entirety of the known chemistry. The opportunities offered by this dataset alongside the next frontiers will be discussed. Much less developed is the exploration of polymeric systems, the second area covered here. The generation of homogeneous datasets (computational and experimental) is in its infancy and, for the same reason, there are major opportunities to shape the development of polymer informatics. Our group managed to develop computational workflows able to explore thousands of polymeric systems but many issues related to system representation, invariants, physical and chemical descriptor remains unsolved.
- 14.30-15.15 (on zoom)
- Greg McColm (Mathematical Crystallography, University of South Florida, US)
- Title. Combinatorial and Topological Equivalence of Representations of Crystals.
- Abstract. Crystals and subperiodic materials (e.g., rods and layers) may be represented by pointsets, directed or undirected graphs, tilings or complexes, or other geometric, topological, or combinatorial structures. Even disregarding issues of numerical discrepancies arising from the environment and instruments, the problem of determining if two representations of the same kind are equivalent can be computationally challenging. We outline a voltage approach to the problem, focusing on graphs, and encounter the most notorious open problem in computer science. Joint work with Nataša Jonoska and Mile Krajcevsk.
- 15.30-16.15 (in person and on zoom)
- Katerina Vriza (GSK, UK)
- Title. Closing the loop between digital models and autonomous experimentation in chemical discovery
- Abstract. Chemical discovery is often limited by fragmented data, heterogeneous instrumentation, and weak coupling between computation and experiment. In this talk, I will present our efforts for closing that loop by integrating multimodal data pipelines, predictive models, and AI agents with automated laboratory workflows. Using examples from real chemical R&D settings, I will highlight the conditions required for these approaches to work in practice: robust data foundations, reliable orchestration across software and hardware, explicit treatment of uncertainty, and meaningful human oversight at critical decision points. The key message is that the impact of AI in discovery depends less on standalone models than on trustworthy closed-loop systems that can continuously learn from, and act on, real-world experiments in both academic and industrial environments.
- 17.00-18.00 (in person)
- Vitaliy Kurlin (Materials Innovation Factory, Liverpool, UK)
- Location: Elizabeth Gidney Room, top floor in the Liverpool Guild of Students
- For catering purposes, please register to attend this inaugural lecture for free.
- Title. Geometric Data Science (inaugural lecture organised by the CSI school).
- Abstract. The emerging area of Geometric Data Science studies real data objects under practical equivalences [1]. The key example is a point cloud under isometry or rigid motion. In a Euclidean space, if given points are ordered, their isometry class is uniquely determined by the matrix of pairwise distances. A naive extension to m unordered points requires exponentially many matrices depending on m! permutations. The case of m=3 points was settled 2000+ years ago due to the side-side-side theorem in school geometry. However, even for m=4 points in the plane, there are infinitely many 4-point clouds that are indistinguishable by 6 pairwise distances. The talk will describe a simple invariant that finished the case of 4 points [2] and a complete invariant with Lipschitz continuous metrics that resolved the exponential challenge in polynomial time in the number of points for any fixed Euclidean dimension [3].
- [1] O.Anosova, V.Kurlin. Geometric Data Science book (arxiv:2512.05040), the latest version at https://kurlin.org/Geometric-Data-Science-book.pdf.
- [2] D.Widdowson, V.Kurlin. Resolving the data ambiguity for periodic crystals. NeurIPS 2022, v.35, p.24625-24638. Extended version in SIAM J Applied Mathematics, v.86 (3), p.898-918 (2026).
- [3] D.Widdowson, V.Kurlin. Recognizing rigid patterns of unlabeled point clouds by complete and continuous isometry invariants with no false negatives and no false positives, CVPR 2023. Extended in arxiv:2303.14161 and MATCH 2026.
- 18.00-19.00 reception (organised by the CSI school)
Back to Top of this page | Back to MACSMIN | Back to Home page
Wednesday 27 May (the MIF boardroom)
- 09.00-09.45 (on zoom) Video (43 min) Slides
- John R. Helliwell (University of Manchester, UK)
- Title. Precision and Accuracy in Biological Crystallography, Diffraction, Scattering, Microscopies, and Spectroscopies.
- Abstract. The talk is based on the recent book, which provides the first comprehensive treatment of uncertainties in the structures determined by all relevant experimental methods; includes an improved set of database resources, governed by data deposition policies set by experts in the field; and highlights the need for new and improved methods which in turn will lead to important developments in research and equipment manufacturing.
- 10.00-10.45 (in person) Video (40 min)
- Thérèse Malliavin (University of Lorraine, France)
- Title. Influence of Stereochemistry in a Local Approach for Calculating Protein Conformations.
- Abstract. Prediction of conformations for intrinsically disordered proteins is mostly based on local structure information, at the contrary of folded protein structure prediction, in which the use of local conformational information is coupled with long-range distance restraints obtained from sequence alignments. The interval Branch-and-Prune algorithm permits to calculate protein conformations using only local conformations knowledge. In that case, the efficiency of calculation is directly related to the knowledge of stereochemistry (bond angles and ω dihedral angles) along the protein sequence and is particularly sensitive to the variations of the dihedral angles ω (da Rocha et al, 2024). The impact of stereochemistry variations is particularly strong in the case of protein topologies defined from numerous long-range restraints, as in the case of protein of β secondary structures. Machine learning methods for classifying ω values would thus be very useful in the frame of conformation prediction for disordered proteins. The prediction of dihedral angles φ and ψ from their sequences has been widely studied in the literature, but the prediction of ω angles has been less explored because it presents more difficulties (Keck et al, 2025). In particular, these angles vary in narrower intervals than the other dihedral angles of the backbone. In addition, random variations of ω angles are produced by the fit of the protein conformations to the X-ray electronic density due to the noise present in the density. Using the interval Branch-and-Prune, we intend to induce a correlation between backbone angles, to improve the efficiency of prediction by Support Vector Machine (SVM), while minimizing the backbone drift induced by modify the angle values. The effects of angle modifications on the prediction of ω angles within sets of homologous proteins and within CATH families of structures, will be presented.
-
References:
(da Rocha et al, 2024) da Rocha W, Liberti L, Mucherino A, Malliavin TE. Influence of Stereochemistry in a Local Approach for Calculating Protein Conformations. J Chem Inf Model. 2024 Dec 9;64(23):8999-9008. doi: 10.1021/acs.jcim.4c01232.
(Keck et al, 2025) Keck U, Guermeur Y, Malliavin TE. Effectiveness of Support Vector Machines for the prediction of dihedral angles ω in proteins. Nancy Computational Structural Biology (NCSB) meeting, 19 June 2025, Nancy.
- 11.00-11.45 (in person) Video (40 min)
- Alexandre de Brevern (Bioinformatics, Université Paris Cité, France).
- Title. Protein Blocks as a discrete geometric model for protein conformational spaces.
- Abstract. Protein structures are commonly described by secondary-structure elements such as helices, strands, and loops, but this coarse representation is often insufficient to capture fine local deformations, transient conformers, and disorder-related rearrangements. Structural alphabets provide a more systematic discretization of protein geometry. Among them, the Protein Block (PB) alphabet, composed of 16 local backbone prototypes, offers a compact and precise encoding of three-dimensional protein conformations.
We present a PB-based framework for representing proteins and molecular dynamics trajectories as symbolic paths in a discrete conformational space. By encoding overlapping backbone fragments into PB sequences, static structures and trajectory ensembles can be transformed into objects that are both geometrically meaningful and computationally tractable. This discrete formalism enables residue-level analysis of local state occupancy, transition frequencies, conformational heterogeneity, persistence, and deformability. It also supports direct comparison of proteins, structural ensembles, and dynamical behaviours using sequence-like or graph-based methods.
Such a representation is particularly useful for identifying flexible hinges, transient local states, weakly structured segments, and disorder-prone regions that are often difficult to characterize using global descriptors such as RMSD or averaged coordinates alone. More generally, PBs provide an effective bridge between continuous protein geometry and discrete algorithmic analysis, allowing the study of conformational landscapes in terms of symbolic dynamics, local transitions, and geometric state organization.
This framework is well suited to the exploration of protein conformational spaces, the analysis of flexibility and disorder, and the characterization of disease-associated local structural changes. It therefore offers a mathematically manageable representation for linking geometry, dynamics, and function in computational structural biology.
- 12.15-13.30 lunch (covered for speakers by the organisers)
- 14.00-14.45 (in person and on zoom)
- Olga Anosova (Data Science Theory and Applications group, Liverpool, UK)
- Title. Data Science exposed thousands of exact duplicates in high-profile structural databases.
- Abstract. The talk discusses several databases of solid crystalline materials and protein structures, such as Google's GNoME AI and the Protein Data Bank (PDB), where every entry is given by dozens or hundreds of atomic positions in Euclidean or fractional coordinates with respect to a unit cell.
Unfortunately, all such cells discontinuously change under almost any noise. This discontinuity of cell-based representations has been resolved by a hierarchy of geometric invariants starting with the ultra-fast vectorial and matrix descriptors, which distinguish all non-duplicate structures in the world’s largest databases of experimental materials, by using only atomic centers without chemical elements, and finishing with the complete isoset invariant and Lipschitz continuous metrics that separate all periodic sets of points under isometry.
Though independent experiments and non-trivial simulations are highly unlikely to produce identical numerical outputs, the Geometric Data Science approach in [1-3] revealed thousands of exact duplicates with all x,y,z coordinates identical. Detecting exact and near-duplicates is essential to avoid biases in AI and ML models that use this data as an input.
[1] O. Anosova, V. Kurlin, M. Senechal. The importance of definitions in crystallography. IUCrJ, v.11(4), p.453-463 (2024).
[2] O. Anosova, A. Gorelov, W. Jeffcott, Z. Jiang, V. Kurlin. Complete and bi-continuous invariant of protein backbones under rigid motion. MATCH, v.94(1), p.97-134 (2025).
[3] A. Wlodawer, Z. Dauter, P. Rubach, W. Minor, M. Jaskolski, Z. Jiang, W. Jeffcott, O. Anosova, V. Kurlin. Duplicate entries in the Protein Data Bank: how to detect and handle them. Acta Cryst D, v.81, 170-180 (2025).
- 15.00-15.45 (in person)
- Alexei Lisitsa (Computer Science, University of Liverpool, UK)
- Title. Automated reasoning for knots and knotted structures with possible applications to proteins.
- Abstract. In this talk, I will present applications of automated reasoning to problems in knot theory and related knotted structures. Fundamental algorithmic tasks—such as unknot detection (i.e., deciding whether a given knot is trivial) and knot equivalence - can be approached by translating these problems into algebraic invariants, including variants of quandles.These translations enable the use of automated reasoning techniques, such as first-order theorem proving, countermodel finding, and constraint-based methods including SAT solving. I will outline how these approaches provide effective tools for analysing knot properties and, in some cases, for certifying non-equivalence. Potential applications include knot detection and analysis in biological and chemical contexts, such as proteins and polymers, where knotted structures naturally arise.
- 16.00-16.45 (in person) Video (39 min)
- Paweł Rubach (Warsaw School of Economics, Poland)
- Title. Knot or not? Identifying unknotted proteins in knotted families with sequence-based Machine Learning model.
- Abstract. Knotted proteins, although scarce, are crucial structural components of certain protein families, and their roles continue to be a topic of intense research. Capitalizing on the vast collection of protein structure predictions offered by AlphaFold (AF), this study computationally examines the entire UniProt database to create a robust dataset of knotted and unknotted proteins. Utilizing this dataset, we develop a machine learning (ML) model capable of accurately predicting the presence of knots in protein structures solely from their amino acid sequences. We tested the model's capabilities on 100 proteins whose structures had not yet been predicted by AF and found agreement with our local prediction in 92% cases. From the point of view of structural biology, we found that all potentially knotted proteins predicted by AF can be classified only into 17 families. This allows us to discover the presence of unknotted proteins in families with a highly conserved knot. We found only three new protein families: UCH, DUF4253, and DUF2254, that contain both knotted and unknotted proteins, and demonstrate that deletions within the knot core could potentially account for the observed unknotted (trivial) topology. Finally, we have shown that in the majority of knotted families (11 out of 15), the knotted topology is strictly conserved in functional proteins with very low sequence similarity. We have conclusively demonstrated that proteins AF predicts as unknotted are structurally accurate in their unknotted configurations. However, these proteins often represent nonfunctional fragments, lacking significant portions of the knot core (amino acid sequence).
- 18.00-21.00 dinner (covered for speakers by the organisers)
Back to Top of this page | Back to MACSMIN | Back to Home page
Thursday 28 May (CMD Seminar room)
- 09.00-09.45 (in person) Video (47 min)
- Saulius Gražulis (Crystallography Open Database, Vilnus, Lithuania)
- Title. Towards the catalogue of entangled nets and knots in the COD and other open databases.
- Abstract. Topological features of materials, such as entanglements of infinite molecular nets, knots in molecules (catenanes), and sterically restricted compounds (rotaksanes) are interesting by themselves and important to understand and design materials with new unexpected properties. As molecular databases grow, we should benefit from efficient procedure that allow us to find such features in structural databases. Unfortunately, there are no straightforward algorithms that would permit efficient search of topological properties in large collections of molecular descriptors.
We present here an algorithm, an implementation, and a database of entangled nets detected in the Crystallography Open Database (COD), the largest currently available open-access collection of crystal structures. I'll also discuss our future plans for determining nets and links in the COD, mention problems that we encounter on the way, and offer some of our thoughts on how to solve them for discussion.
- 10.00-10.45 (in person)
- Wolfgang Hornfeck (Institute of Physics, Czech Academy of Sciences)
- Title. Arithmetic and algebraic patterns in crystal structures.
- Abstract. The classification of crystal structures is traditionally based on the mathematical concept of groups (space, plane, point, etc.). While group theory in general provides a most powerful framework of describing the symmetry of periodically long-range ordered atomic arrangements in three dimensions, it is also well known that it does not encompass all kind of symmetries which might be present. For instance, it cannot account for the full point group symmetry of molecules exhibiting a non-crystallographic rotational symmetry incompatible with lattice translations. Yet, there are still more, and more subtle, patterns hidden in the atomic arrangements, and their coordinate representations, which might be described by other mathematical concepts from the fields of arithmetic, algebra, and geometry, respectively. In my talk I will give a number of examples: (i) similar sublattice structures described by multiplicative congruential generators; (ii) two- and three-dimensional permutation structures; (iii) Z module twin quasilattices. These examples for what one might call quantitative crystal structure descriptors will be discussed in relation to actual crystal structures of intermetallic compounds. Moreover, my talk will address general questions of randomness and order, spatial uniformity, as well as structural complexity in the algorithmic sense of Kolmogorov.
Hornfeck, W. (2012). Acta Cryst. A 68, 167-180; Hornfeck, W. (2013). Acta Cryst. A 69, 355-364; Hornfeck, W. (2025). Struct. Chem. 36, 2007-2019.
- 11.00-11.45 (on zoom)
- Yury Elkin (Data Science Theory and Applications group, Liverpool, UK)
- Title. A geometric map of chemical elements based on molecular databases QM9 and GEOM.
- Abstract. Detecting chemical elements from only geometric data is important for structure determination, because X-ray diffraction or other experimental tools usually identify only positions of atomic centres without chemical types. We introduce simple geometric invariants that detect elements with high accuracy in molecules from the QM9 and GEOM databases. All chemical elements in these databases occupy disjoint regions in low-dimensional spaces of invariant values.
- 12.00-13.30 lunch (covered for speakers by the organisers)
- 13.30-14.15 (in person and on zoom) Video (44 min)
- Daniel Widdowson (Data Science Theory and Applications group, Liverpool, UK)
- Title. The Crystal Geomap visualises materials databases in real time.
- Abstract. Our rigorously justified invariants of crystals give rise to a continuous space containing all crystals, where the proximity of two crystals does not depend on a choice of unit cell and motif, but whether the two structures can be closely matched atom for atom by isometry [1]. This led us to develop software to visualise this space and be an interface to ultra-fast comparisons of crystals enabled by our invariants [2]. In this talk we will explore unusual “features” of crystal databases such as the ICSD made visible by our depictions of crystal space, examples of nearly identical crystals represented with completely different cells and motifs [3], and a live example of detection of all geometric (near-)duplicates in the ICSD, a calculation which was computationally intractable by existing methods.
[1] O.Anosova, V.Kurlin, M.Senechal. The importance of definitions in crystallography. IUCrJ, v.11(4), p.453-463 (2024).
[2] D.Widdowson, V.Kurlin. Continuous invariant-based maps of the Cambridge Structural Database. Crystal Growth & Design, v.24(13), p.5627–5636 (2024).
[3] D.Widdowson, V.Kurlin. Geographic-style maps with a local novelty distance help navigate the materials space. Scientific Reports, v.15, 27588 (2025).
- 14.30-14.50 (on zoom) Video (18 min)
- Surya Majumder (Data Science Theory and Applications group, Liverpool, UK)
- Title. Continuous invariant-based asymmetries of periodic crystals quantify deviations from higher symmetry.
- Abstract. Ideal symmetry is known to break down under almost any noise. One measure of asymmetry in a periodic crystal is the relative multiplicity Z′ of geometrically nonequivalent units. However, Z′ discontinuously changes under almost any displacement of atoms, which can arbitrarily scale up a primitive cell. This discontinuity was recently resolved by a hierarchy of invariant descriptors that continuously change under all small perturbations. We introduce a Continuous Invariant-based Asymmetry (CIA) to quantify (in physically meaningful Angstroms) the deviation of a periodic crystal from a higher symmetry form. Our experiments on several Crystal Structure Prediction datasets show that about a half of simulated crystals have high values of CIA, while all experimental structures in these datasets have CIA = 0. On another hand, many crystals with high values Z′ in the Cambridge Structural Database (CSD) turned out to be close to more symmetric forms with Z′≤1 due to low values of CIAs. The talk is based on the paper in IUCrJ 2026.
- 15.00-15.45 (in person)
- Richard Catlow FRS (Materials Science, University College London, UK)
- Title. Crystal and Nano-Structure Prediction: Achievements and Opportunities.
- Abstract. Crystal structure prediction is long standing and major challenge in the field of computational condensed matter science1,2; while the modelling and prediction of nano-particle structures is an increasingly active and topical field. This lecture will aim to summarise both the main developments in the field over recent years and the current opportunities. We will discuss energy landscape search procedures , and consider recent developments relating to the use of quantum computing and of AI/ML techniques, focussing on applications to inorganic materials and metal nano-particles. We will conclude by considering future developments in the field.
(1) Woodley SM and Catlow R, Nature Materials 7, 937, (2008)
(2) Woodley SM, Day GM and Catlow R, Phil Trans Roy Soc A, 378, 20190600, (2020)
- 16.00-16.45 (on zoom)
- Simon Billinge (Materials Science, University of California Santa Barbara, US)
- Title. Symmetry in a Hierarchical World.
- Abstract. When an object fills space uniformly, such as a crystal, we can describe its symmetry uniquely by assigning it its space group. We can think of this symmetry as being a binary quantity: the object either has the symmetry or it does not. The group consists of a set of symmetry operations, each of which is satisfied by the object. If any of the elements is not satisfied, the space-group changes. However, we can ask a question: what if we introduce a small defect in the structure, for example, by introducing a vacancy on a single atomic site. Strictly speaking, the periodic symmetry is broken and we can't even use space groups any more. But the change is infinitely small, so practically speaking, we still want to be able to apply group theory in this case, it can still be a useful tool for us. In practice, we recover by doing some averaging and recovering a periodic signal and a deviatoric correction and we can apply the space-group symmetry to the periodic part. But this introduces the concept of "an approximately broken symmetry" or a "continuously changing" symmetry. And it allows us to ask questions such as "If I introduce 'such and such' a defect, which of the symmetry operations will be strongly broken and which will be weakly broken?". Working on the basis that such a concept could be useful to science, we have explored ways to quantify symmetry breaking. I will introduce these ideas in this talk. I introduce a method that uses a statistical approach based on the Jensen-Shannon divergence. The measure is calculated by comparing the transformed atomic density function with its original after applying the symmetry transformation. I will introduce the measure and and show some tests for various cases including finite size crystallites (where the surfaces break the crystallographic symmetry), atomic displacements from high symmetry positions, and collective motions of atoms due to rotations of rigid octahedra. The approach provides a powerful tool for assessing local symmetry breaking and offers new insights that can help researchers understand how different structural distortions affect different symmetry operations.
- 18.00-21.00 dinner (covered for speakers by the organisers)
 At the dinner from left to right:
Wolfgang Hornfeck,
Saulius Gražulis,
Vitaliy Kurlin,
Alex Wlodawer,
Alexandre de Brevern,
Paweł Rubach,
Ziqiu Jiang (Henry),
Dan Widdowson,
Richard Catlow FRS,
Thérèse Malliavin,
Olga Anosova.
At the dinner from left to right:
Wolfgang Hornfeck,
Saulius Gražulis,
Vitaliy Kurlin,
Alex Wlodawer,
Alexandre de Brevern,
Paweł Rubach,
Ziqiu Jiang (Henry),
Dan Widdowson,
Richard Catlow FRS,
Thérèse Malliavin,
Olga Anosova.
Back to Top of this page | Back to MACSMIN | Back to Home page
Friday 29 May (the MIF boardroom)
- 09.00-09.45 (in person)
- Alex Wlodawer (Structural Biology, National Cancer Institute, US)
- Title. Community efforts to improve the contents of the Protein Data Bank, a crucial resource for structural biology.
- Abstract. For over half a century, the Protein Data Bank (PDB) has been accumulating the experimental structures of biological macromolecules, becoming a crucial resource created by structural biologists for the use of the whole biological and biomedical community and beyond. With over quarter of a million experimentally determined structure models, it has also served to provide training data for the remarkably successful artificial intelligence-based prediction of protein structures. Implementation by the PDB curators of validation tools has led throughout the years to vast improvement in the quality of PDB deposits, but occasional inaccuracies, mistakes, and errors of different severity are still able to sneak into the database despite tight scrutiny. Several such problems were identified during campaigns that analyzed the quality of structure models of selected protein families, such as β-lactamases, SARS-CoV-2 proteins, cAMP-dependent protein kinases, or L-asparaginases. In many cases, the problems could be attributed to depositors' negligence, lack of training or supervision, or cutting corners by ignoring the warnings in the PDB Validation Reports. Postulates directed to the PDB included elimination of exact duplicates, development of better mechanisms for linking series of model updates, finding a better home for group depositions resulting from fragment screening campaigns, and prominent display of CAVEAT records to warn human and machine users of potential problems in the PDB deposits. Other identified general problems included the lack of modeled solvent in many high-resolution protein crystal structures. Such problem-tracking efforts ultimately aim to alert the PDB and, if possible, suggest solutions to make this crucial resource even better.
- 10.00-10.45 (in person and on zoom)
- Ziqiu Jiang (Data Science Theory and Applications group, Liverpool, UK)
- Title. Atomic clashes in the Protein Data Bank (PDB).
- Abstract. The validation of 3D structures in the PDB involves several scores, most importantly, the clashscore counting an average number of atomic clashes per 1000 atoms. For presumably distant (at least non-bonded) atoms, a clash between atoms is considered serious, if the inter-atomic distance is smaller than the sum of the van der Waals radii of these atoms, minus a tolerance threshold of 0.4A. Less than 1% entries (not pure DNA/RNA) in the PDB have a clashscore of 0. The talk will discuss the recently found atomic clashes in the majority of these entries, some of which are easy to correct.
- 11.00-11.20 (on zoom)
- Jack Gallimore (Mathematics, University of Leeds, UK)
- Title. Geometry based method for identifying hydrogen bonds and classifying helices.
- Abstract. Understanding secondary structure in proteins is fundamental to structural biology, yet current algorithms rely on fixed geometric cutoffs and hand tuned rules, making them both discrete and hard to interpret. We propose a geometry-based framework that identifies hydrogen bonding using statistically justified continuous regions in geometric space, without rigid thresholds. By analyzing 110M+ residues across 707K+ geometrically valid chains in the Protein Data Bank, we identify small regions in the geometric distribution of interatomic distances and angles that correspond to hydrogen bonding. These regions, rather than fixed cutoffs, naturally reflect the spatial variation observed in experimental data and enable us to detect secondary structure, particularly α-helices. This method offers a more interpretable and flexible approach to structural annotation, with potential to unify helix classification, including, including 310 and π-helices.
- 11.30-11.50 (in person and on zoom)
- Gabriel Newton (Data Science Theory and Applications group, Liverpool, UK)
- Title. ProtNRD: A 3D Ramachandran Dashboard for Exploring Neighbouring Residue Influences on Backbone Geometry.
- Abstract. The Protein Neighbouring Residue Dashboard (ProtNRD) addresses a critical gap in structural biology education: the lack of tools for visualising how neighbouring residues influence backbone geometry. ProtNRD is an intuitive, open-source, web-based 3D visualisation platform operating across two complementary modes — a pairwise filtering mode analysing residue influences across ±4 step windows, and a 3-mer analysis mode isolating specific triplet conformations. Through targeted amino acid filters, users can systematically investigate how local sequence composition constrains or permits particular backbone geometries, with support for experimental Protein Data Bank data (X-ray resolution ≤2Å) and a modular data pipeline with the potential to extend to custom sources such as synthetic AI-generated structures. The latest release is available at https://github.com/moracore/protNRD/releases/latest.
- 12.00-14.00 lunch (covered for speakers by the organisers)
Back to Top of this page | Back to MACSMIN | Back to Home page
Travel information: venue, accommodation, trains, and flights
- All talks in person will be in the ground floor boardroom in the Materials Innovation Factory (MIF), Liverpool, UK. Address: 51 Oxford street, building 807 in the grid cell F5 on the campus map. The building has a secure entrance, so we will let the reception know about MACSMIN participants. The MIF is 15 min on foot from the Liverpool Lime Street station.
- If you contact us in advance, we can help with booking hotels. One option is the Liner hotel in a quiet street close to the Liverpool Lime Street main rail station. Explore other good hotels and attractions on the website visit Liverpool.
- The city has the Liverpool John Lennon airport with convenient buses to the centre. The larger Manchester airport has the train station with direct 90-min trains to the Liverpool Lime Street station. Check flights to nearby airports at Skyscanner.
Back to Top of this page | Back to MACSMIN | Back to Home page
